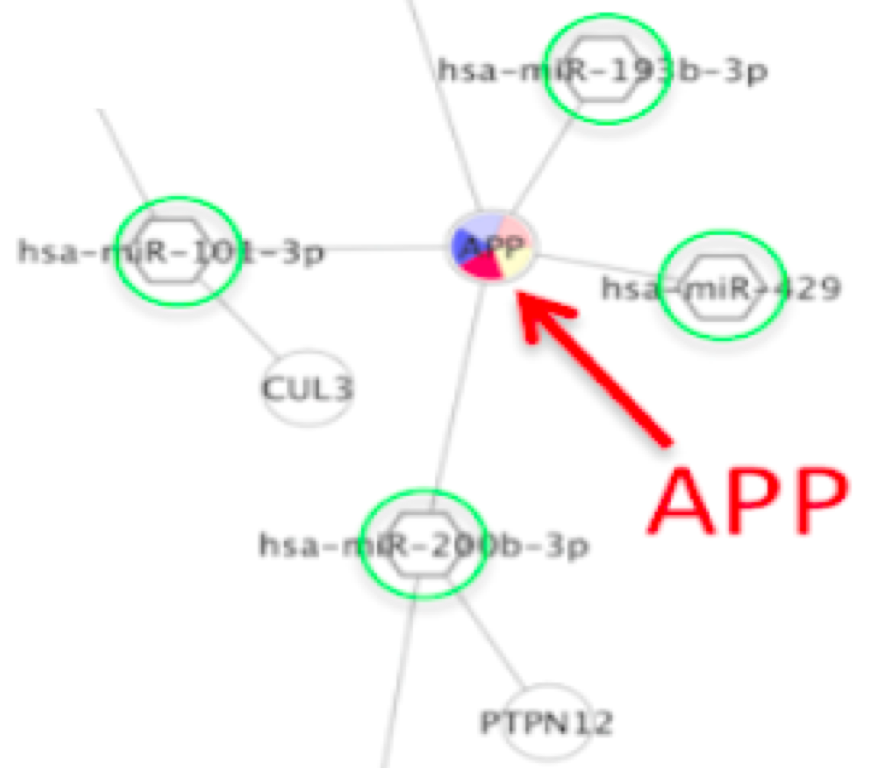
(a) Differentially co-expressed module networks with the DiffCorr package. Specifically, it has 15842 nodes and 49812 edges. Figure 1.7 Module network visualization with the DiffCorr package and Cytoscape. Several case studies of Cytoscape plug-ins are surveyed, including a search for interaction pathways correlating with changes in gene expression, a study of protein complexes involved in cellular recovery to DNA damage, inference of a combined physical/functional interaction network for Halobacterium, and an interface to detailed stochastic/kinetic gene regulatory models. Which is the best way for visualizing a big network in Cytoscape I'm using Cytoscape for visualizing my transcriptional network, but it is huge. The Core is extensible through a straightforward plug-in architecture, allowing rapid development of additional computational analyses and features. Interactive network visualization in Python and Dash, powered by Cytoscape.js - GitHub - plotly/dash-cytoscape: Interactive network visualization in Python. Cytoscape is not too hard to use, but it won’t make much sense unless you have a sense of some basic network analysis vocabulary and concepts. Cytoscape's software Core provides basic functionality to layout and query the network to visually integrate the network with expression profiles, phenotypes, and other molecular states and to link the network to databases of functional annotations. Cytoscapeis a tool for viewing and analyzing networks(meaning, in this case, any group of entities that are connected in some way). Information to browse the database and use its. The ggpubr package was used to perform Spearman correlation analysis on these hub genes. Designed to find pathways and network patterns related to cancer and other types of diseases. Although applicable to any system of molecular components and interactions, Cytoscape is most powerful when used in conjunction with large databases of protein-protein, protein-DNA, and genetic interactions that are increasingly available for humans and model organisms. Then, Cytoscape plugin-Analyze Network was used to screen the significant modules with the degree >3 in the PPI network, and hub genes from CS-DEGs were obtained. 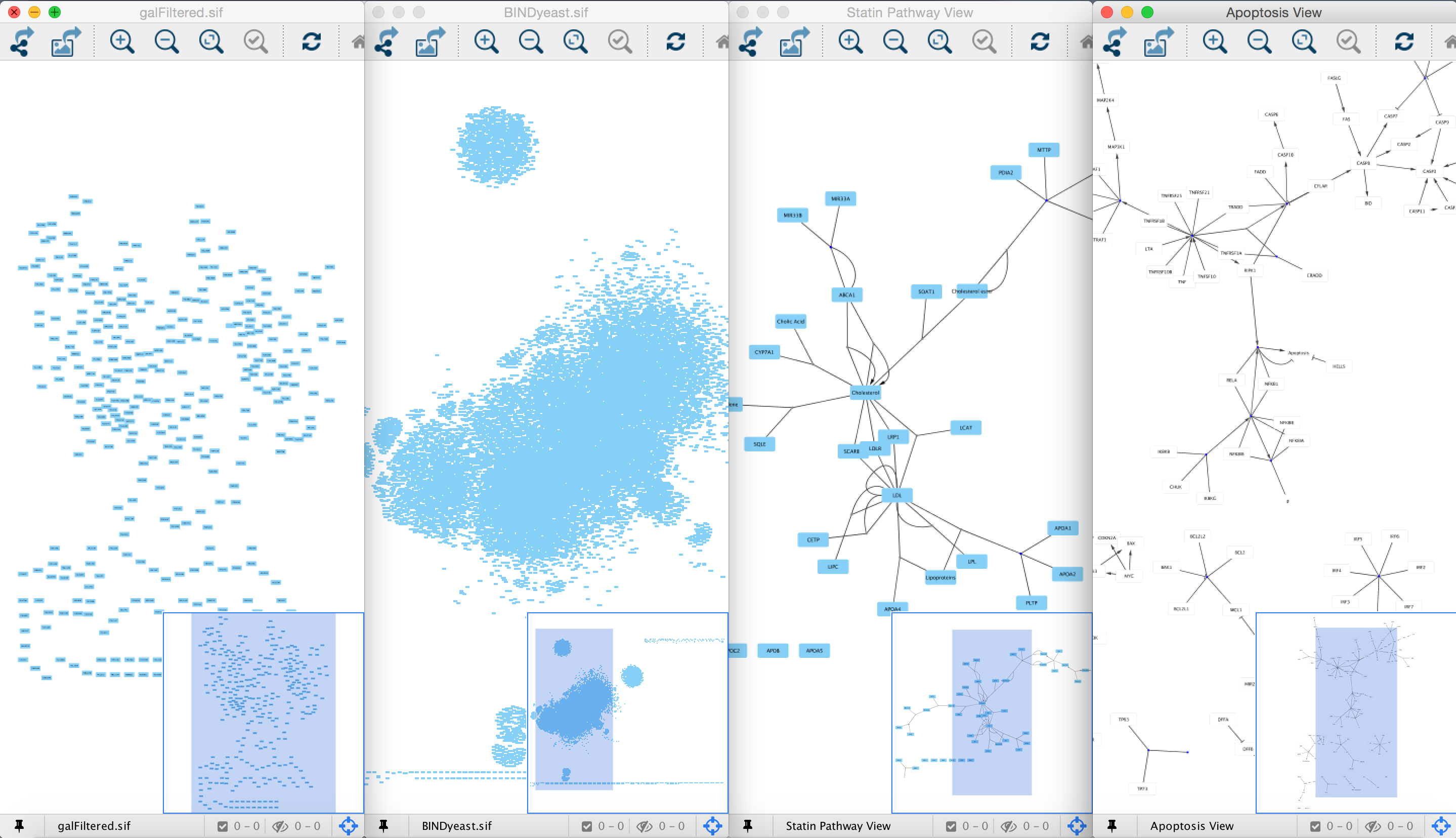
Cytoscape is an open source software project for integrating biomolecular interaction networks with high-throughput expression data and other molecular states into a unified conceptual framework.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
